Research Project: CRISPR Genome Engineering
A brief History of Genome Engineering
DNA breaks in most eukaryotes are repaired by either the error prone mechanism non-homologous end-joining (NHEJ) or precisely repaired by homologous recombination (HR). Typically HR is initiated in S- or G2-phase of the cell cycle when a sister chromatid is available to provide a donor homology template for repair by HR (Figure 1A). This property of HR was used in the early 1980s to both repair and insert genes in mammalian cells and eventually mouse embryos to generate transgenic mice, and the modern era of gene engineering was born (Capecchi., 2005). These early seminal studies came from the laboratories of Drs. Oliver Smithies, Mario Capecchi and Martin Evens, who pioneered homologous recombination-mediated gene editing in mouse embryos and would later share the 2007 Nobel Prize in Physiology or Medicine “for their discoveries of principles for introducing specific gene modifications in mice by the use of embryonic stem cells“.
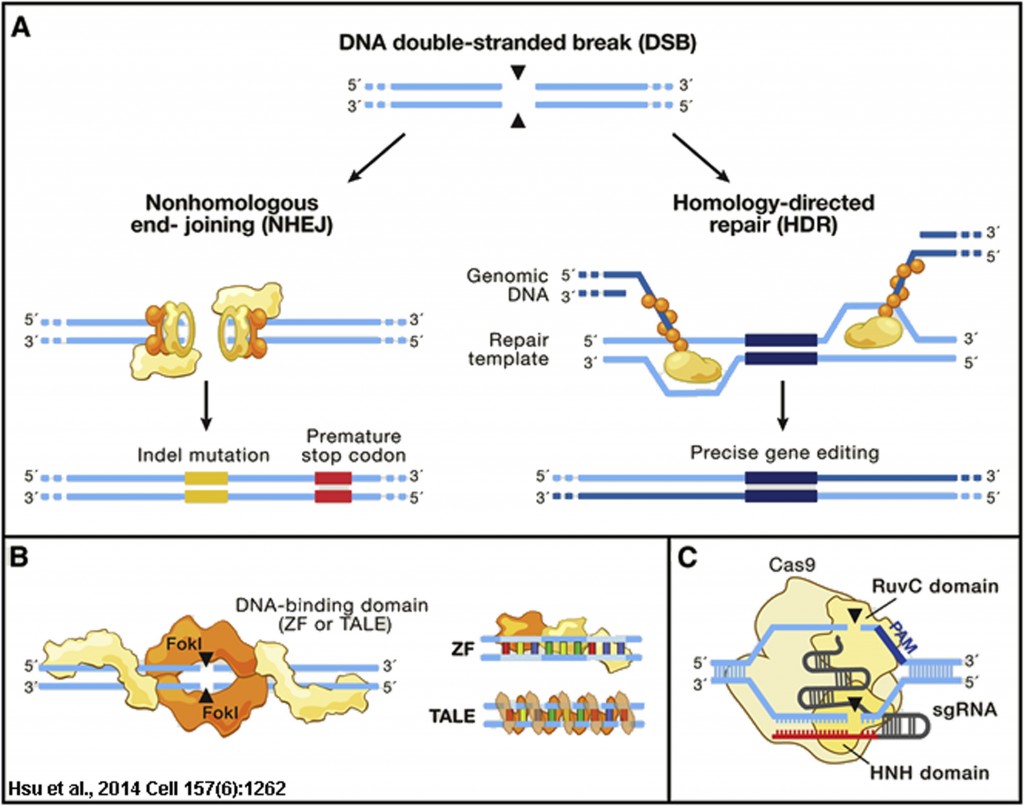
Figure 1: Genome Editing Technologies Exploit Endogenous DNA Repair Machinery
The next advancements in gene targeting using homologous recombination would come from Dr. Maria Jasin and colleagues in th late 1990s, who employed the yeast homing endonuclease I-SceI to generate site-specific breaks in chromatin carrying an ectopic copy of the 18-bp rare restriction site for the SceI enzyme. Cutting with the SceI enzyme could enhance HR up to 500 fold at the target locus, and these studies would foreshadow the use of designer site-specific endonucleases to enhance gene editing (Rouet et al., 1994; Smih et al., 1995). These designer endonucleases included the zinc finger nucleases (ZFNs) and the Transcription Activator-Like Effector Nucleases (TALENs), which combined the DNA binding specificity of zinc fingers or TALE transcription factors from plants with the FokI endonuclease DNA cutting activity (Figure 1B). However, both ZFNs and TALENs had issues with limited targeting abilities, were expensive and time consuming to construct and test, and/or were too large or complex for genome-wide screening or viral delivery as would be required for gene therapies. This would all change with the discover and adaptation of the bacterial innate immune system of Streptococcus pyogenes, known as CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) and encoding the Cas9 RNA-directed endonuclease (Figure 1C), to genome engineering (Hsu et al., 2014).
The story of the CRISPR-Cas9 system, began with the discovery of a cluster of 29 bp repeats down stream of the iap gene locus in E. coli by Nakata and colleagues in 1987 (Ishino et al., 1987), which would later be discovered to represent a unique form of clustered repeats found in >40% of bacterial species and later collectively referred to as Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) by Jansen and Mojica in 2002 (Jansen et al., 2002). Associated with these repeats were a number of CRISPR-associated genes (Cas) including Cas9, which was shown by Moineau and colleagues to be an RNA-directed endonuclease in 2010 (Garneau et al., 2010). The Cas9 protein of the Type II CRISPR system of S. pyogenes most widely used for genome engineering binds two RNAs encoded within the CRISPR repeats, the crRNA (CASCADE complex for type I; Cmr or Csm RAMP complexes for type III) and the tracrRNA (transactivating CRISPR RNA)(Deltcheva et al., 2011). The crRNA and the tracrRNA hybridize promoting cleavage by the host RNASe III into a double-stranded hybrid crRNA/tracrRNA that when associated with the Cas9 protein acts to guide the cleavage of the target DNA 3 nucleotides 5′ to the protospacer-adjacent motif (PAM); a sequence of 3 nucleotides adjacent to the binding site of the gRNA within the target DNA, e.g. NGG for S. pyogenes where N = any nucleotide. The crRNA/TracrRNA can be combined into a single chimeric guide RNA (gRNA) and Charpentier and Doudna demonstrated that this single gRNA could target site specific cleavage of DNA (Jinek et al., 2012). The CRISPR-Cas9 system was finally applied to genome editing by homology directed repair (HDR) in mammalian cells by the groups of George Church and Feng Zhang in papers published in the same issue of Science in 2013 (Cong et al., 2013 Science; Mali et al., 2013 Science). Since the seminal work of the Charpentier/Doudna and Church/Zhang groups described above, the number of papers citing the use of CRISPR is doubling roughly every year, with more than 1400 publications just in 2015. The CRISPR-Cas9 system is extremely versatile and being applied to not only gene knock-out studies but to gene therapies for muscular dystrophy (Long et al., 2014), gene regulation through the use of enzymaticaly dead dCas9 fused to transcriptional activators and repressors (reviewed in Dominguez et al., 2016), epigenetic regulation through dCas9 targeting of histone modifying enzymes (Hilton et al., 2015), as well as chromosome tagging studies (Chen et al., 2013).
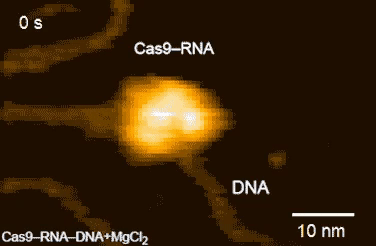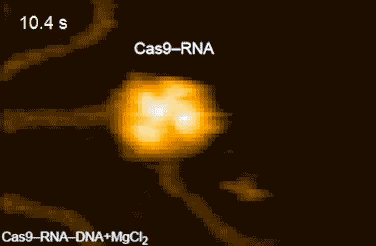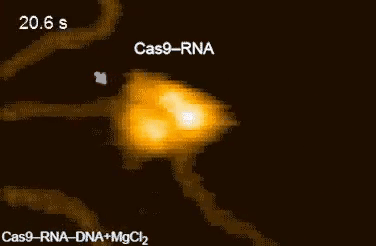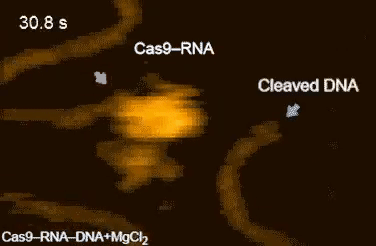
| Movie 1. CRISPR in Action. The Cas9/sgRNA complex cleaving DNA visualized by high speed atomic force microscopy (From Shibata et al., 2017 Nature Communications). |
 |
CRISPR Genome Editing – simple, effective, and highly versatile
In our laboratory we have been working to enhance CRISPR-mediated genome editing through the use of small molecules, as well as developing tools for measuring the efficiency of homology directed repair using CRISPR. For example, we developed a screen based on the insertion of the bright EGFP variant gene Clover into the lamin A/C locus (LMNA) by CRISPR-mediated HDR (Pinder et al., 2015) and this system was used in collaboration with the group of Dr. Daniel Durocher to identify the molecular basis for the inhibition of HR in non-dividing cells, an advance that could open the way to gene therapy of diseases such as muscular dystrophies that affect non-dividing muscle cells (Orthwein et al., 2015). We continue to develop novel assays and tools using CRISPR technology for the study of DNA repair and cancer. Our current projects include:
- Enhancing the efficiency of CRISPR-mediated Homology Directed Repair (HDR) genetically and through novel small molecules
- Applying CRISPR-HDR to the identification and characterization of nuclear domain associated proteins
- Development of novel transgenic and gene-modified zebrafish for the study of cancer
Other Resources
- CRISPR gene editing: Why we need Slow Science
- Moratorium on Human Genome Editing: Time to Get It Right
- Why we are not ready for genetically designed babies
- He Jiankui: A Sorry Tale of High-Stakes Science
References
Capecchi MR. (2005) “Gene targeting in mice: functional analysis of the mammalian genome for the twenty-first century“. Nat Rev Genet 6(6):507-12.
Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, Park J, Blackburn EH, Weissman JS, Qi LS, Huang B. (2013) “Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system.” Cell 155(7):1479-91.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W,Marraffini LA, Zhang F. (2013) “Multiplex genome engineering using CRISPR/Cas systems“.Science. 339(6121):819-23
Deltcheva, E., Chylinski, K., Sharma, C.M., Gonzales, K., Chao, Y., Pirzada, Z.A., Eckert, M.R., Vogel, J., and Charpentier, E. (2011). “CRISPR RNA maturation
by trans-encoded small RNA and host factor RNase III“. Nature 471(7340):602–607
Dominguez AA, Lim WA, Qi LS. (2016) “Beyond editing: repurposing CRISPR-Cas9 for precision genome regulation and interrogation.” Nat Rev Mol Cell Biol. 17(1):5-15.
Garneau, J.E., Dupuis, M.E., Villion, M., Romero, D.A., Barrangou, R., Boyaval, P., Fremaux, C., Horvath, P., Magada´ n, A.H., and Moineau, S. (2010). “The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA“. Nature 468(7320):67–71
Hilton IB, D’Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE, Gersbach CA. (2015) “Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates
genes from promoters and enhancers.” Nat Biotechnol. 33(5):510-7.
Hsu PD, Lander ES, Zhang F. (2014) “Development and applications of CRISPR-Cas9 for genome engineering“. Cell 157(6):1262-78.
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. (1987) “Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product.” J Bacteriol. 169(12):5429-33.
Jansen, R., Embden, J.D., Gaastra, W., and Schouls, L.M. (2002). “Identification of genes that are associated with DNA repeats in prokaryotes“. Mol. Microbiol.
43, 1565–1575.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. (2012) “A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity“.Science. 337(6096):816-21.
Long C, McAnally JR, Shelton JM, Mireault AA, Bassel-Duby R, Olson EN. (2014) “Prevention of muscular dystrophy in mice by CRISPR/Cas9-mediated editing of
germline DNA. Science. 345(6201):1184-8.
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM. (2013) “RNA-guided human genome engineering via Cas9“. Science 339(6121):823-6.
Orthwein, A., S.M. Noordermeer, M.D. Wilson, S. Landry, R.I. Enchev, A. Sherker, M. Munro, J. Pinder, J. Salsman G. Dellaire, B. Xia, M. Peter, and D. Durocher. (2015) “Induction of homologous recombination in G1 cells” Nature 528(7582):422-426
Pinder, J., J. Salsman, and G. Dellaire. (2015). “Nuclear domain “knock-in” screen for the evaluation and identification of small molecule enhancers of CRISPR-based genome editing” Nucleic Acids Research 43(19):9379-9
Rouet P, Smih F, Jasin M. (1994) “Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells.” Proc Natl Acad Sci U S A.
91(13):6064-8
Smih F, Rouet P, Romanienko PJ, Jasin M. (1995) “Double-strand breaks at the target locus stimulate gene targeting in embryonic stem cells“. Nucleic Acids Res. 23(24):5012-9